Statistics
Contents
Anti-CRISPRdb contains the available protein sequences, DNA sequences, coding regions, source organisms, taxonomy, virulence, and protein interactors and the corresponding three-dimensional structures.
Now version 2.2 contains more information: inhibitory stages and mechanisms, inhibitory ability for Acr-Cas/Acr-CRISPR systems, the estimation of Acr neighbors. Additionally, we also proposed CUBRank, a parameter to measure if a gene is in GIs.Data summary
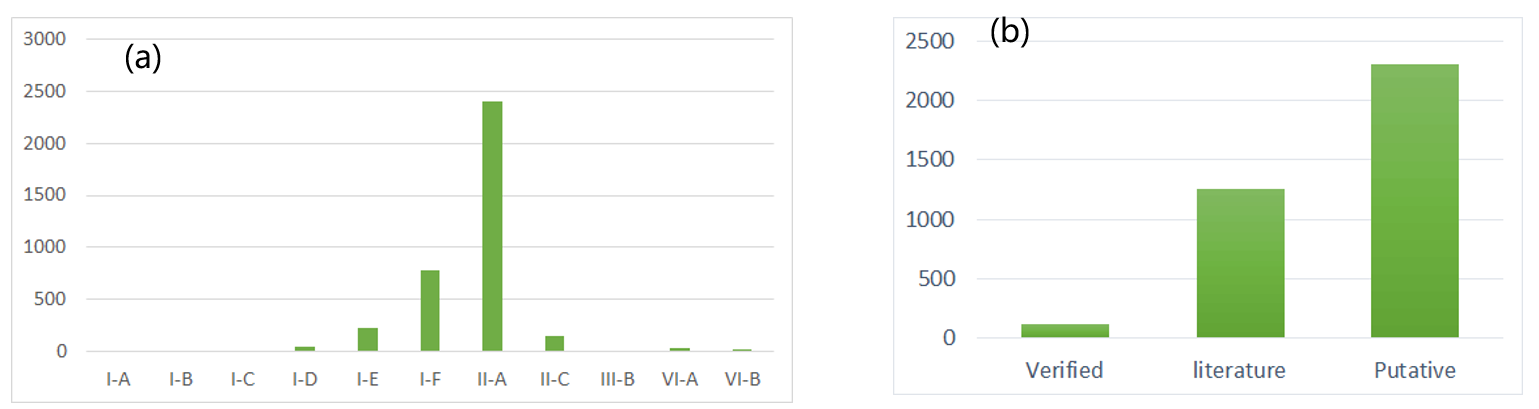
Verified Acr entries: 122, Acrs from PLiterature: 1253, putative Acrs: 2304 (figure 1b), which can be splited into 91 families among the records. Those families can inhibit a wide range of CRISPR-Cas system including I-A~I-F, II-A, II-C, VIA, VAB and III, which distribution is illustrated in figure 1a
Five inhibiting types including prevent spacer acquisition, guid RNA loading, DNA binding, DNA cleavage and degradate cA4 in signaling pathway.

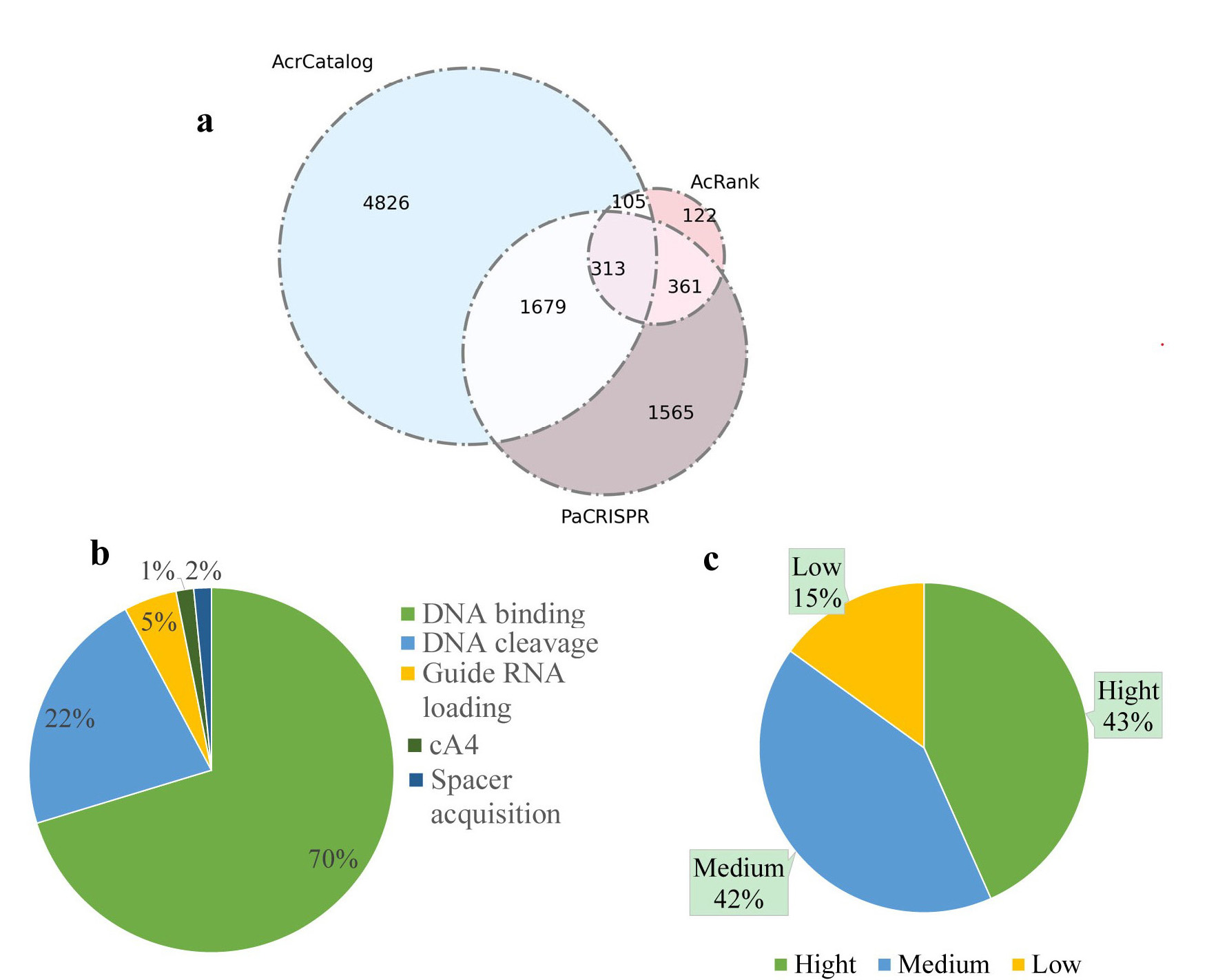
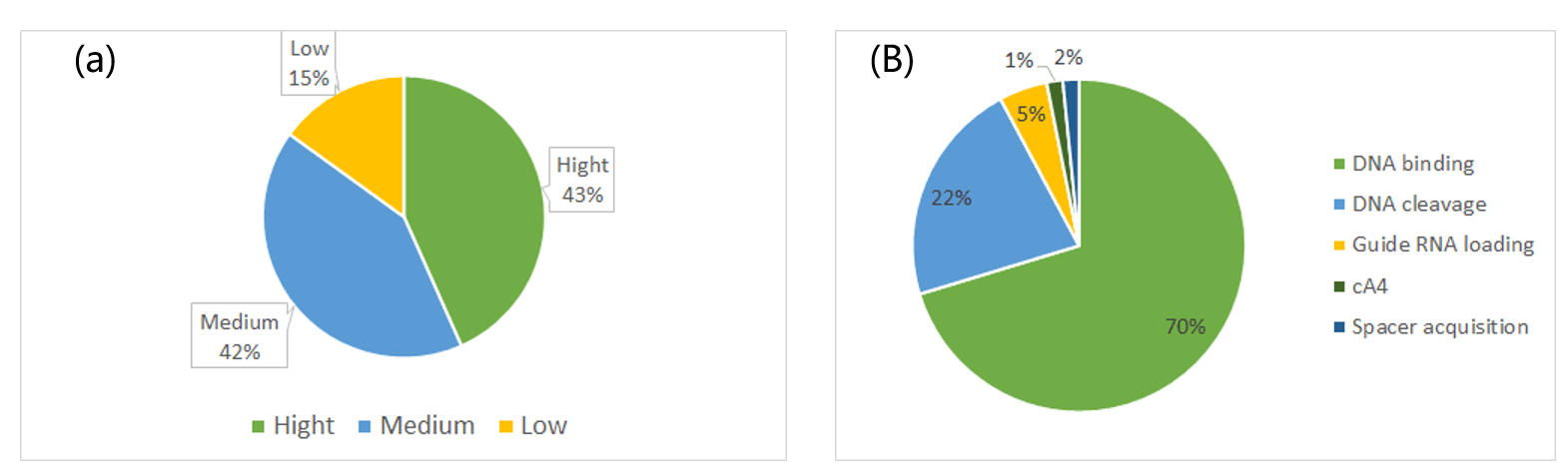
According references, we divided the inhibitory strength into three levels: high, medium, low. Among them 43% Acr-Cas or Acrs-CRISPR-Cas pairs have high blocking strength. The remaining 57% pairs have medium and low records (a). There are five inhibitory mechanisms: inhibit DNA binding (70%), which is the majority part; inhibit DNA cleavage (22%); inhibit guide RNA loading (5%); other 3% including inhibit spacer acquisition and inhibit cA4 signal pathway (b).
There are totally 13,040 unique neighbors in our newly updated database, including neighbors belonging to our putative Acrs. Among these proteins, 6,923 unique proteins showed similarities in the AcrCatalog database when we adopted an E-value less than or equal to 0.01 and an identity higher than or equal to 35% as cutoffs. 901 unique proteins were ranked in the top 10 according to AcRank; and 3,918 unique proteins were predict-ed to be Acrs by PaCRISPR. A total of 2,458 proteins (1,679+313+105+361) were predicted to be potential Acrs by more than two of the three methods, among which 313 proteins were predicted by all three predicted algorithms